Computational Analysis on The Development of New Technetium-99m-labeled Pentapeptide for Cancer Molecular Imaging Targeting Integrin α5β1
Abstract
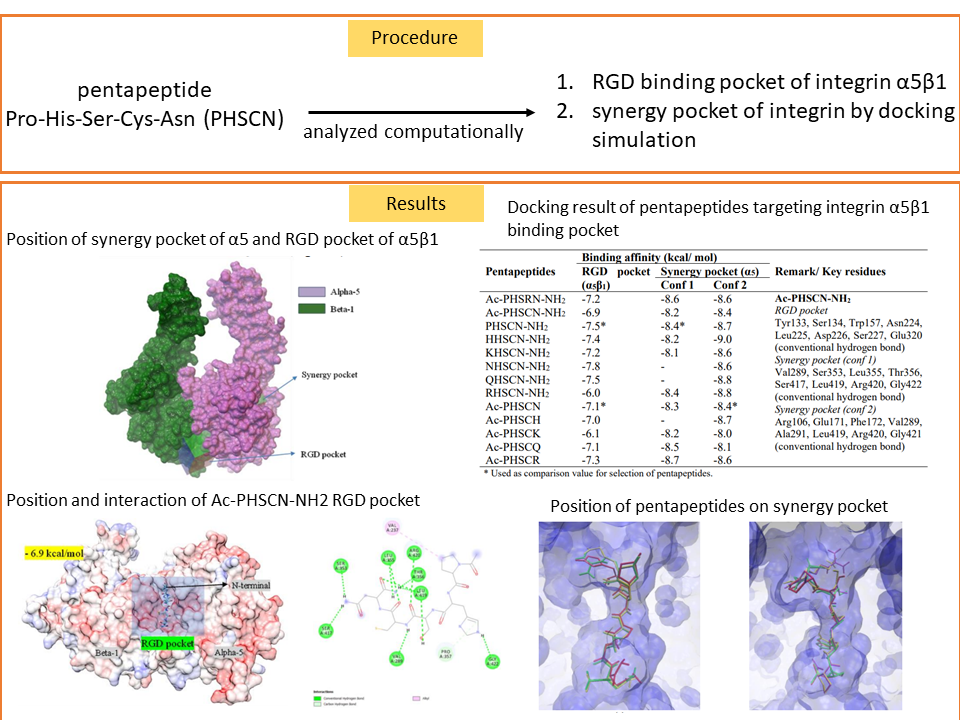
Cancer continues to be a major leading cause of death despite huge efforts dedicated to developing anticancer drugs. Radiopeptide is currently used for targeted therapy and diagnosis of cancer. The structure selection of new radiopeptide should be determined to minimize the decrease in binding affinity because of metal radioisotope and its chelator. In this research, the interaction of radiopeptides based on technetium metal on RGD binding pocket of integrin α5β1 and synergy pocket of integrin α5 was analyzed by molecular docking simulation using Autodock Vina and Autodock 4. Pentapeptide Pro-His-Ser-Cys-Asn (PHSCN) has two possible conformations to interact with integrin α5 which is predicted could be labeled in two possible positions. Even though the results showed that the binding value of radiolabeled compounds was lower than PHSCN’s, some radiolabeled compounds in this simulation might have biological activity. The use of HYNIC-EDDA as a chelator produces better value than DTPA and MAS3. Radiolabeling procedures in the N-terminal position of the peptide are preferable and have higher affinity than C-terminal modification. Further laboratory experiments are required to confirm the activity of EDDA-Tc-HYNIC-PHSCN-NH2 on the integrin α5β1 receptor.
References
[1] H. Sung, J. Ferlay, R. L. Siegel, M. Laversanne, I. Soerjomataram, A. Jemal and F. Bray, CA. Cancer J. Clin., 2021, 17 (3), 209–249
[2] M. Jamous, U. Haberkorn, and W. Mier, Molecules, 2013, 18, 3, 3379–3409
[3] W. Chiangjong, S. Chutipongtanate, and S. Hongeng, Int. J. Oncol., 2020, 57 (3), 678–696
[4] F. Schaffner, A. M. Ray, and M. Dontenwill, Cancers, 2013, 5 (1), 27–47
[5] A. Sureshkumar, B. Hansen, and D. Ersahin, Seminar in Ultrasound CT and MRI, 2019, 41 (1), 10–19
[6] A. Boschi, L. Uccelli, and P. Martini, Appl. Sci., 2019, 9 (12), 2526
[7] M. Nagae, S. Re, E. Mihara, T. Nogi, Y. Sugita, and J. Takagi, J. Cell Biol., 2012, 197 (1), 131–140
[8] O. Trott and A. J. Olson, J. Comput. Chem., 2010, 31, 455-461
[9] G. M. Morris, R. Huey, W. Lindstrom, M. F. Sanner, R.K. Belew, D. S. Goodsell, and A. J. Olson, J. Comput. Chem., 2009, 30 (16), 2785–2791
[10] A. Baldi, Syst. Rev. Pharm., 2010, 1 (1), 99-105
[11] S. P. Leelananda and S. Lindert, Beilstein J. Org. Chem., 2016, 12 (1), 2694–2718
[12] Y. Feng and M. Mrksich, Biochem., 2004, 43 (50), 15811–15821
[13] D. Kozakov, L. E. Grove, D. R. Hall. T. Bohnuud, S. Mottarella, L. Luo, B. Xia, D. Beglov, and S. Vajda, Nat. Protoc., 2016, 10 (5), 733–755
[14] J. Giglio and A. Rey, Inorg., 2019, 7 (11), 128
[15] I. T. Ibrahim, M. El-Tawoosy, and H. M. Talaat, ISRN Pharmaceutics, 2011, 578570, 1-6
Refbacks
- There are currently no refbacks.

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.